INTRODUCTION
The indiscriminate use of antibiotics in human and animal medicine, as well as for prophylaxis and growth promotion in animal husbandry, threatens to reduce the effectiveness of these fundamental drugs. Antibiotic-resistant bacteria, although occurring naturally, are also released into the environment, where they may out compete sensitive bacteria due to the presence of antibiotics and other chemical contaminants that are also released into the environment. The resistance genes in these bacteria can then be transferred to other pathogenic and non-pathogenic bacteria, thereby increasing the environmental reservoir of resistant bacteria and genetic resistance determinants.
Escherichia coli is a commensal bacterium that inhabits the intestines of humans and other animals and is often used to indicate environmental fecal contamination. Some E. coli are pathogens that can cause urinary tract, gastrointestinal or nosocomial infections. Antibiotic-resistance in environmental E. coli has also been proposed as an indicator to monitor the extent of antibiotic resistance in the environment 1.
The bacterial load of wastewater discharged into natural water bodies is significantly reduced by treatment plants. However, these plants may also promote the spread of antibiotic-resistant bacteria and resistance genes by providing favorable conditions for increasing the relative abundance of resistant bacteria and the horizontal transfer of the genes conferring this resistance 2. Although there are few treatment plants in Venezuela and much of the wastewater is discharged directly into the environment, there is a wastewater treatment plant in the state capital Maracaibo, located in the “El Tablazo” Petrochemical Complex of the Miranda municipality of Zulia state. The plant was designed so that the petrochemical industry could reuse some of its effluent water while the rest would be discharged into the giant Lake Maracaibo. There have been very few studies on antibiotic-resistance in bacteria isolated from raw or treated wastewater in Venezuela, but such studies represent essential surveillance measures to assess the extent of antibiotic-resistance in the environment and plan corrective strategies. Accordingly, we set out to perform a phenotypic and molecular study of antibiotic resistance in E. coli isolates from the wastewater treatment plant mentioned above.
MATERIALS AND METHODS
E. coli was isolated from water samples of the “El Tablazo” treatment plant (Miranda Municipality, Zulia State) collected from May to October 2012 for a microbiological quality evaluation. The system includes a pre-treatment to remove solids followed by absorption, biological oxidation, and a first chlorine injection. The water is then transported to the plant at “El Tablazo” and subjected to physical and biological treatment based on reactors where dissolved organic matter is removed, followed by a secondary settling. Then a first effluent is discharged into Lake Maracaibo. Another portion of the water to be used by the petrochemical complex is treated with a flocculant and chlorine.
Sampling sites were four different sections of the treatment plant: pre-treated influent (Site1); after physical processing (Site 2); after biological processing (effluent to the Maracaibo lake, Site 3); and the chlorine disinfection point (Site 4). There were six water samples from Site 1 and Site 4, four from Site 2, and five from Site 3.
The water samples were collected in sterile bottles and processed according to the procedures described in the Standard Methods for examination of Water and Wastewater to determine coliform by the fermentation technique 3. Samples showing growth in EC broth were streaked onto EMB agar to select typical E. coli colonies, which were sub-cultured in nutrient agar tubes for transport. Re-isolation was performed on McConkey agar, and colonies were cultured in LB broth and then stored in 20% glycerol at -80°C. All assays were performed on the bacteria regrown from the frozen stocks.
Biochemical identification and antibiotic susceptibility testing
Bacterial isolates were first identified with the following biochemical tests: TSI, indole-motility, methyl red, Voges Proskauer, citrate, and urea.
Resistance was assessed with the Kirby Bauer disc diffusion method, using commercial discs with the following antibiotics: tetracycline 30 µg (TE), ampicillin 10 µg (AMP), ampicillin-sulbactam 10/10 µg (SAM), sulfisoxazole 250 µg (SF), chloramphenicol 30 µg (C), trimethoprim 5 µg (W), trimethoprimsulfamethoxazole 1.25 µg /23.75 µg (SXT), ciprofloxacin 5 µg (CIP), aztreonam 30 µg (ATM) and imipenem 10 µg. E. coli ATCC 25922 was used as an antibiotic-susceptible control strain. The results were interpreted according to CLSI guidelines 4.
PCR amplification
PCR was used to confirm the bacteria as E. coli, determine phylogroups, and detect the presence of resistance genes and the integrase gene intI1. All PCR reactions were performed on boiled bacterial lysates, using Taq DNA polymerase with ThermoPol buffer (NEB), following the manufacturer’s instructions, using previously reported specific primers, some of which were modified as indicated below. PCR was performed to amplify genes conferring resistance to tetracycline (tetA and tetB) 5, sulfisoxazole (sul1, sul2, and sul3) 6-8, chloramphenicol (floR and cat) 6, ampicillin (blaTEM) 9 and trimethoprim (dfrA12 and dfrA7&17) 10,11. The detection of the intI1 integrase was with primers described by Moura12. In contrast while the phylogroup identification was performed using the quadruplex plus group C specific PCR described by Clermont et al. 13 Negative controls without template DNA were included in each PCR assay. A subset of the PCR products were confirmed by DNA sequencing (Macrogen, Korea) and used as positive controls for the detection of resistance genes and intI1, as well as the determination of phylogroups.
The reactions were performed with previously reported primers and conditions or with the following variations: Sul1-R. 5’-TGATCTAACCCTCGGTCTCT-3’ temperature of annealing (Ta) 56°C, blaTEM-F 5’-GCATACACTATTCTCAGAATGA-3’ blaTEM-R 5’-CTCACCGGCTCCAGATTTAT-3’ Ta 56°C, dfr7&17-F 5’-CATTTGACTCTCTATGGGTGTTC TT-3’ Ta 58°C.
To avoid analyzing duplicate resistant isolates, REP-PCR was performed on isolates showing the same phenotype and genotype, using the REP1 and REP2 primers as previously described 14. E. coli-specific PCR was performed using primers to amplify rrs (75F and 619R) or gad15,16, with E. coli XL1-blue as a positive control.
Transformation and conjugation assays
Transferability of the detected antibiotic resistance genes was assessed by heat shock transformation using transformation competent E. coli DH5α as the recipient strain and plasmid DNA obtained by alkaline lysis from all isolates resistant to tetracycline, ampicillin, chloramphenicol, or trimethoprim in which PCR had detected a resistance gene. Transformants were selected on LB agar plates containing either carbenicillin (50 µg/mL), ampicillin (32 µg/mL), tetracycline (30 µg/mL), chloramphenicol (30 µg/mL), or trimethoprim (20 µg/mL) as appropriate. Phenotypic resistance was confirmed for each transformant, and the presence of plasmid DNA and the relevant resistance genes were verified.
In some cases, the capacity for conjugation was assessed in liquid medium using E. coli J62-2 as the recipient strain. Selection was performed on LB agar plates supplemented with tetracycline (30 µg/mL) and rifampicin (50 µg/mL). Transconjugants were confirmed by phenotypic resistance, amplification of the same resistance gene detected in the donor, and ERIC-PCR with primers described by Versalovic et al. 14.
RESULTS
Here, we characterized thirty-six isolates identified as E. coli with biochemical tests and identified as E. coli with biochemical tests and PCR amplification of rrs or gad. These isolates originated from the four sampling sites: 13 from Site 1, four from Site 2, nine from Site 3, and ten from Site 4.
Resistance phenotypes
As shown in Fig. 1, the highest frequency of resistance was to tetracycline and the lowest to ciprofloxacin and ampicillinsulbactam, with intermediate prevalences of resistance to the other antibiotics tested, including ampicillin.
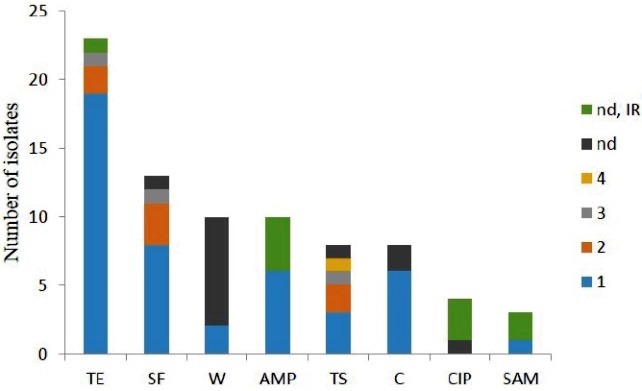
Fig. 1 Phenotypic and genotypic resistance per antibiotic. The numbers of phenotypically resistant isolates with corresponding genotypes (for full resistance) are indicated. Resistance genotypes are numbered from 1 to 4 according to the detection frequency from highest to lowest. 1: tetA (TE); sul2 (SF and TS); dfrA12 (W); floR (C); or blaTEM (AMP and SAM). 2: tetB (TE); or sul3 (SF and TS). 3: tetA-tetB (TE); sul1 (SF); sul1-dfrA12(TS). 4: sul3-dfrA12 (TS). nd: Resistance genotype not determined, IR: intermediate resistance. TE, tetracycline; SF, Sulfisoxazole; W, trimethoprim; C, Chloramphenicol; TS, Trimethoprim-sulfamethoxazole; AMP, ampicillin; SAM, ampicillin-sulbactam; CIP, ciprofloxacin.
Twenty-four isolates (66.6%) were fully resistant to at least one antibiotic, corresponding to 7/13, 2/4, 7/9, and 8/10 isolates from sampling points 1 to 4, respectively (Table 1). Five of these 24 isolates (20.8%) also showed intermediate resistance to one or two additional antibiotics (two AMP-CIP, one CIP, and two SAM). Among the 12 remaining isolates, three had only intermediate resistance (two AMP and one TE), and nine (9/36, 25%) were fully sensitive to all antibiotics tested. There were also seven isolates (7/24, 29.1%) fully resistant to three or more antibiotics of different classes and thus multi-drug resistant or MDR. Two of these originated from each of sampling Sites 1, 3, and 4, and one isolate was from Site 2 (see Table 1). All isolates were sensitive to aztreonam and imipenem.
Table 1 Phenotypic and genotypic profiles of the bacterial isolates.
| Sampling site | Resistance phenotype (FR/IR) | Resistance genotype | Phylogroup |
|---|---|---|---|
| 1 | TE | tetB | B1 |
| 1 | TE-SF-W-TS | tetA-sul2 | C |
| 1 | TE-SF-W-TS | tetA-sul2 | C |
| 1 | TE-SF-W-TS/amp-cip | tetA-sul2 | C |
| 1 | TE-SF- TS-C/amp-cip | tetA-sul2-floR | A |
| 1 | TE-SF-C/ cip | tetA-sul2-floR | A |
| 1 | TE-C | tetA-floR | A |
| 1 | amp | nd | B1 |
| 1 | S | na | C |
| 1 | S | na | C |
| 1 | S | na | C |
| 1 | S | na | C |
| 1 | S | na | B2 |
| 2 | TE | tetA | C |
| 2 | TE-SF-W-TS-AMP-C/sam | tetA-sul1-blaTEM-floR-dfrA12-intI1 | C |
| 2 | S | na | A |
| 2 | S | na | A |
| 3 | TE | tetA | B1 |
| 3 | TE-W | tetA | C |
| 3 | TE-SF-W- TS-AMP-C-CIP/sam | tetA-sul3-blaTEM-intI1 | C |
| 3 | TE-SF-W-TS- AMP-SAM | tetA-sul3-blaTEM-dfrA12-intI1 | C |
| 3 | TE-AMP | tetB-blaTEM | C |
| 3 | AMP-C | blaTEM- floR | B1 |
| 3 | AMP-W | blaTEM | C |
| 3 | te | nd | C |
| 3 | S | na | A |
| 4 | TE | tetA | C |
| 4 | TE | tetA | A |
| 4 | TE-SF | tetA | B1 |
| 4 | TE-SF | tetA-sul2 | A |
| 4 | TE-SF | tetA-sul2 | C |
| 4 | TE-SF | tetA-sul2 | B1 |
| 4 | TE-SF-W-TS-C | tetA-sul3-intI1 | C |
| 4 | TE- W-C | tetA- tetB- flor-intI1 | A |
| 4 | amp | nd | C |
| 4 | S | na | C |
FR: Fully resistant, IR: intermediate resistance (lowercase), S: sensitive, nd: not determined, na: not apply. The abbreviations for the antibiotics are the same as in Fig. 1.
Resistance genes
The genotypic resistance profiles and genes detected are described in Table 1. The most frequently detected genes were tetA (20/22) and sul2 (8/12), while all isolates fully resistant to ampicillin contained blaTEM (Fig. 1). The floR gene was detected in most (6/8) of the chloramphenicol-resistant isolates, none of which contained the cat gene. The only amplified determinant associated with trimethoprim resistance was dfrA12, which was detected in just two of the ten trimethoprim-resistant isolates.
Of the seven MDR-resistant isolates, the intIl gene was amplified from 5, representing 21% of the resistant isolates and 13.9% of total isolates (Table 1). In one of the isolates in which the intIl gene was detected, amplification and sequencing with primers specific for conserved segments of class 1 integrons detected dfrA12 and aadA2, which encodes an aminoglycoside adenyl transferase conferring streptomycin resistance.
Transfer of resistance determinants
Transformants were obtained from the isolated plasmid DNA of 20/24 resistant isolates. Most of the transformants (18/20) were recovered on media with tetracycline, while only 1/20 were recovered on media with chloramphenicol and 1/20 on media with trimethoprim. However, the major part of the genes conferring resistance to sulfisoxazole and chloramphenicol were co-transferred with the tetA gene (Table 2). Despite repeated attempts, no transformants were obtained from plasmid DNA isolated from the remaining four resistant isolates.
Table 2 Antibiotic resistance profiles of transformants and transconjugants.
| Genotype | Phenotype | Number of transformants (N=20) | Number of transconjugants (N=2) |
|---|---|---|---|
| tetA-sul2 | TE-SF | 8 | - |
| tetA-sul3 | TE-SF | 1 | - |
| tetA | TE | 6 | 2 |
| tetA-floR | TE-C | 3 | - |
| tetA- sul3- dfrA12 | TE-SF-W | 1 | - |
| flor- blaTEM | C-AMP | 1 | - |
The tetA gene was transferred from almost all isolates in which it was detected (19/20). The blaTEM gene was transferred from 1/6 of the isolates in which it was detected, the sul genes were transferred from 10/12 isolates (8 sul2 and 2 sul3), floR from 4/6 and dfrA12 from 1/2 isolates containing this gene. The tetB gene was not transferred under the conditions employed. The resistance genes most frequently detected in our isolates, tetA and sul2, hybridized with plasmid DNA isolated from most (18/20), or all isolates (8/8), respectively, in which the genes had been detected by PCR, confirming that they were carried on plasmids (not shown).
The transfer capability of the tetA gene was tested by conjugation experiments with five isolates in which the tetA gene was detected. Transconjugants were obtained from two isolates at the high frequencies of 7.5 x 10-3 y 1.95 x 10-2, demonstrating that the tetA gene was carried on conjugative plasmids at least in these two isolates. One of these five isolates with the tetA gene generated neither transformants nor transconjugants; nevertheless, only scant plasmid DNA could be obtained from this isolate, perhaps because its plasmid was either very large or present at a very low copy numbers.
E. coli phylogroup analysis
Of the 36 bacterial isolates studied, 20 belonged to group phylogenetic C, nine to group A, six to group B1, and one to group B2 (Table 1).
DISCUSSION
In the present work, we analyzed phenotypic antibiotic-resistance, detected corresponding antibiotic-resistance genes, evaluated their transferability, and determined the phylogroups of 36 E. coli isolates obtained from a wastewater treatment plant. Similar to other studies of resistant E. coli isolates from wastewater and treatment plants 17-21, the most frequent resistance encountered was to tetracycline, sulfonamide, trimethoprim, and ampicillin. In contrast, resistance to the carbapenems and ciprofloxacin was infrequent or not detected. Our results differ from a previous phenotypic study of E. coli isolates from stabilization ponds in Maracaibo, Venezuela, that found a higher proportion of resistance for ampicillin (81.25%) followed by tetracycline and trimethoprim 21. In comparison, a study of E. coli isolates from a treatment plant in Cumaná (Venezuela) found 73.3% and 23.3% of ampicillin and tetracycline resistance, respectively. These discrepancies may be due to differences in the treatment systems.
Although tetracycline and sulfonamide, to which we found medium to high frequencies of resistance (22% to 64%), have been, for the most part, replaced by newer agents in human medicine, they are still classified as highly important by the World Health Organization.23 Ampicillin (28% resistance) is still frequently used and considered critically important in human medicine, as are the antibiotics for which we found less frequent resistance (8% to 11%).
The antibiotic resistance we observed is common in E. coli isolated from healthy humans in low and middle-income countries 24, and the antibiotic-resistance in human isolates of E. coli has been correlated with the resistance in E. coli isolated from the local wastewater 25. The resistance patterns may also be affected by selection or adaptation to the specific characteristics of the treatment plant 26. A high prevalence of the tetA, sul2, and blaTEM genes has been previously observed in E. coli isolated from other treatment plants.17,27 However, previous studies have predominantly found the cat gene in chloramphenicol-resistant isolates 17,25, but most of our chloramphenicol resistant isolates carried the floR gene. In contrast, the cat gene was not detected.
We found the intI1 gene in 13.9% of total isolates, similar to a study by Figueira et al. that found the gene in 22.3 % of E. coli isolates from treated wastewater 20. Integrons contribute to the spread of multidrug resistance, and class I integrons are the most important in clinical isolates 28.
Horizontal transmission of plasmids carrying resistance genes is a crucial route for disseminating resistance. Similar to our results, a previous study of E. coli and other fecal coliforms isolated from treatment plants found that most plasmid transformants were resistant to tetracycline, followed by chloramphenicol and trimethoprim 29. The resistance genes tetA, blaTEM, tetB, sul, floR, and dfrA12 can be found on chromosomes, but as we observed, they are frequently carried on plasmids 30-35.
E. coli isolates can be classified into seven main phylogenetic groups: A, B1, B2, C, D, E, and F. Intestinal pathogenic E. coli strains have been associated with groups A, B1, and E, while extra-intestinal strains mainly belong to groups B2 and D36. However, Group C can also include human pathogenic strains, as shown in a study of human isolates from the USA and Europe, in which phylogroup C was associated with uropathogenic E. coli, although B2 and D predominated 37. Jafari et al. found that group C was the most common (21.3%) among Shiga toxin-producing E. coli (STEC) patient isolates 38. It has been suggested that some E. coli phylogroups, particularly groups A and B1 (or non-B2), are more prone to develop antibiotic resistance to traditional antibiotics and fluoroquinolones 39,40. Most previous studies assigning E. coli phylogroups have used the triple PCR method 41 that identifies only four groups: A, B1, B2, and D. Several studies have found group A to predominate in wastewater, followed by D or B120. Group A is also the phylogroup most frequently observed in human commensal isolates, followed by B2 group 42. Researchers employing the quadruplex method plus the group C specific PCR, or in silico typing, have found that phylogroup C is less frequent than other phylogroups in samples from birds, humans, non-human mammals, domestic animals, wild animals, river and lake water43-45. Two studies on strains from wastewater found that group A or B2 predominated, while none or only 1% belonged to group C 17,46.
Therefore, the presence of group C in most of our isolates differs from the findings in similar studies and could be due to geographical location or climate differences, as these factors may influence the distribution of phylogroups 45. It is also possible that previous studies that used only the triple PCR method classified C group isolates as group A.
In conclusion, we observed E. coli isolates resistant to diverse antibiotics, including some clinically essential agents and found that many were associated with transferable genetic determinants and class 1 integrons. Also, in contrast to other studies, many of our isolates belonged to the phylogroup C. The characteristics of the isolates we studied may have been determined by the influent water, the nature of the treatment plant, and environmental conditions, but an evaluation of the contribution of each of these aspects would require a much larger study with many more isolates. Similar studies should be repeated in the same treatment plant and undertaken in other treatment plants in Venezuela, and these studies should be extended to include untreated wastewater and focus on resistance to the newer, currently more commonly used antibiotics.